(best viewed on original website)
Over the last year, my focus has been diverted from exploring analytics, new packages and blogging, to completing my dissertation. With the dissertation now complete and only final edits remaining, I had some spare time to spend on projects that I have been curating throughout the year. One such project that has been in the back of my mind for the last couple of months concern itself with with faster, scalable machine learning. This is where h2o comes in. We have been using h2o in production over the last year with great results. But as the data has grown, so has our need for faster turnaround in the estimation of the models. Currently, 96 core, 750GB RAM machines are not doing the job (Or perhaps I am just impatient). I have a limited amount of time and would like to spend it evaluating the model’s business value, not waiting 12 hours for it to train. This has lead me to two very promising and exciting alternatives:
- Multi Node h2o clusters
- h2o4gpu
Although the multi node h2o cluster approach is something to behold, I wanted to start working with GPUs1!
The rest of this post goes through the pain and suffering of getting to the point where you can train models using the h2o4gpu package in R. It hopefully serves as a resource for users (and future me) who just want to skip the fuss and run a h2o model on a GPU. All of the following setup commands and tests are done on a p2.xlarge machine from amazon. It goes for around $0.38 per hour if you go for a spot request and comes with 12GB of GPU memory, 4 cores and 61GB RAM. I also use Louis Aslett’s amazing AMI as a starting point - ami-09aea2adb48655672.
The steps for getting this to work is (tldr):
- Start
p2.xlargemachine with 100GB HDD - Create virtual environment for python
- Install h2o4gpu into environment
- Install
Rpackage - Do AI
Getting started with h2o4gpu
If you are not familiar with the h2o open source machine learning toolkit, I highly recommend having a look at their website and some of the previous blogs I wrote on how amazing their machine learning suite is - Part I, Part II, Part III.
Apart from the usual h2o api, the team has also been working on h2o4gpu:
H2O4GPU is a collection of GPU solvers by H2O.ai with APIs in Python and R. The Python API builds upon the easy-to-use scikit-learn API and its well-tested CPU-based algorithms. The h2o4gpu R package is a wrapper around the h2o4gpu Python package, and the interface follows standard R conventions for modeling.
Continuing with their vignette, further down it states:
The Python package is a prerequisite for the R package. So first, follow the instructions here to install the h2o4gpu Python package (either at the system level or in a Python virtual envivonment)
And this is where the wheels came off… I decided to install the packages at a system level, which according the the vignette:
… if you installed the h2o4gpu Python package into the main Python installation on your machine…
setting your virtual environment using reticulate::use_virtualenv("/home/ledell/venv/h2o4gpu") is not necessary. Which for the life of me I could not get working due to a couple of frustrations:
- Python2.7 is still default python on ubuntu
- Installing reticulate will most likely install miniconda and keep looking there
This eventually led me to go the venv route, which worked! If you are like me and didnt know how/why python prefers virtual environments, you can read up here. The first thing to do is install the necessary package:
sudo apt install python3-venv python3-pip
sudo update-alternatives --install /usr/bin/python python /usr/bin/python3.6 2
sudo update-alternatives --config python
From here we follow the steps as per the h2o4gpu vignette. First we must make some adjustments in our .bashrc:
export CUDA_HOME=/usr/local/cuda
export LD_LIBRARY_PATH=$LD_LIBRARY_PATH:$CUDA_HOME/lib64/:$CUDA_HOME/lib/:$CUDA_HOME/extras/CUPTI/lib64
Install OpenBlas dev environment:
sudo apt-get install libopenblas-dev pbzip2
We are building the h2o4gpu R package, so it is necessary to install the following dependencies:
sudo apt-get -y install libcurl4-openssl-dev libssl-dev libxml2-dev
Once the installation is done, we need to create and activate the virtual environment:
python3 -m venv h2o4gpu
source h2o4gpu/bin/activate
Next we can download and install the Python wheel file for CUDA:
python -m pip install h2o4gpu
My understanding of wheels are that they resemble the notion of the tidyverse - a collection of packages and dependencies, working together to bring you a service - in my case h2o4gpu2.
Now we can test whether it works!
import h2o4gpu
import numpy as np
X = np.array([[1.,1.], [1.,4.], [1.,0.]])
model = h2o4gpu.KMeans(n_clusters=2,random_state=1234).fit(X)
model.cluster_centers_
# >>> import h2o4gpu
# >>> import numpy as np
# >>>
# >>> X = np.array([[1.,1.], [1.,4.], [1.,0.]])
# >>> model = h2o4gpu.KMeans(n_clusters=2,random_state=1234).fit(X)
# >>> model.cluster_centers_
# array([[1. , 0.5],
# [1. , 4. ]])
# >>>
Back to R
Hopefully all worked, which means you are now well on your way to running models in R on the gpu. Using the AMI discussed above, you can log into the server with:
- username: rstudio
- password: instanceid
The instance id is available on your EC2 dashboard:
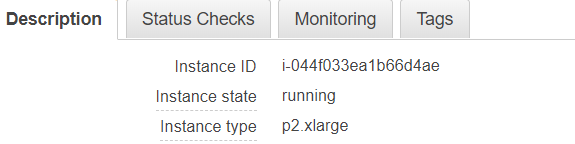
After changing to darkmode you should have the classic AMI screen (restart your session the first time you log in, otherwise packages struggle to install):
To install the development version of the h2o4gpu R package, you can install directly from GitHub as follows:
# install.packages("devtools", Ncpus = 4)
# install.packages("Metrics", Ncpus = 4)
library(devtools)
devtools::install_github("h2oai/h2o4gpu", subdir = "src/interface_r")
Higgs example
To illustrate the package, I will now use the h2o4gpu package to distinguish between signal “1” and background “0”, so this is a binary classification problem. Do note
The h2o4gpu R package does not include a suite of internal model metrics functions, therefore we encourage users to use a third-party model metrics package of their choice. For all the examples below, we will use the Metrics R package. This package has a large number of model metrics functions, all with a very simple, unified API.
First, set up the libraries and the VERY important use of the virtual environment:
library(tidyverse)
library(h2o4gpu)
library(reticulate)
use_virtualenv("/home/ubuntu/h2o4gpu/", required = T)
py_config()
Next, lets load up the example data from Erin LeDell3:
# Setup dataset
train <- read.csv("https://s3.amazonaws.com/erin-data/higgs/higgs_train_10k.csv")
test <- read.csv("https://s3.amazonaws.com/erin-data/higgs/higgs_test_5k.csv")
If you feel like experimenting with the full dataset (11mil observations), you can download it using data.table::fread. Just be aware that you will also need to install R.utils as well:
# install.packages('data.table')
# install.packages('R.utils')
higgs <- data.table::fread("https://archive.ics.uci.edu/ml/machine-learning-databases/00280/HIGGS.csv.gz",
verbose = TRUE,
nThread = 4)
test_index <- sample_frac(tibble(index = 1:nrow(higgs)), 0.7)
train <- higgs[test_index$index,]
test <- higgs[-test_index$index,]
Create train & test sets (column 1 is the response):
x_train <- train[, -1]
y_train <- unlist(train[, 1])
x_test <- test[, -1]
y_test <- unlist(test[, 1])
The final step is to estimate the models, predict on unseen test data and compare the different models. The models are estimated in a two-step fashion: initialize and then fit. This is more common in Python, and they borrow that paradigm in the package.
model_gbc <- h2o4gpu.gradient_boosting_classifier(n_estimators = 10) %>% fit(x_train, y_train)
model_rfc <- h2o4gpu.random_forest_classifier(n_estimators = 10) %>% fit(x_train, y_train)
pred_gbc <- model_gbc %>% predict(x_test, type = "prob")
pred_rfc <- model_rfc %>% predict(x_test, type = "prob")
Metrics::auc(actual = y_test, predicted = pred_gbc[, 2])
Metrics::auc(actual = y_test, predicted = pred_rfc[, 2])
If all went well, your model will be estimated with the GPU. It is quite difficult to tell if the process is working as should if you do not have some kind of monitoring tool. I always use htop for process monitoring, so I found a tool called nvtop. I could only get the older build one as I am not using ubuntu 19.04, which in all honesty sucks. I loved how the picture look:
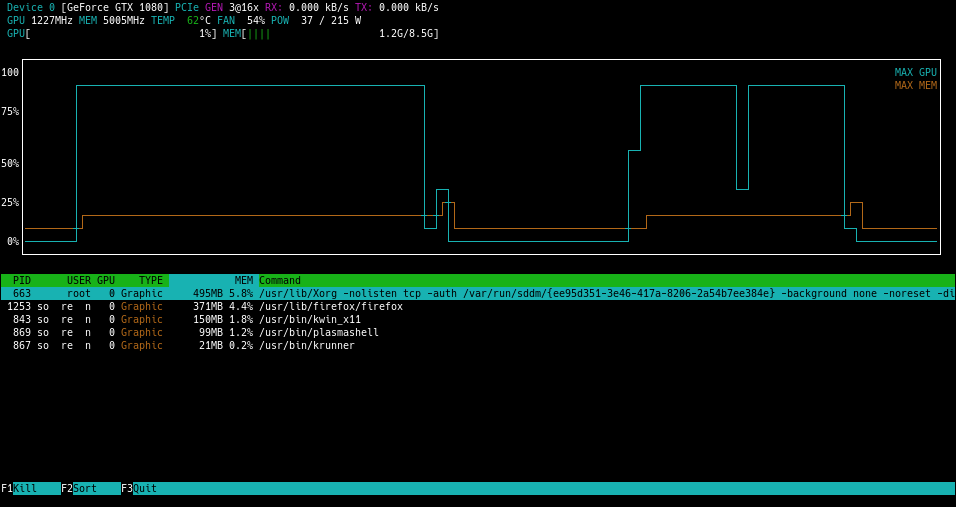
BUT, instead I got this:
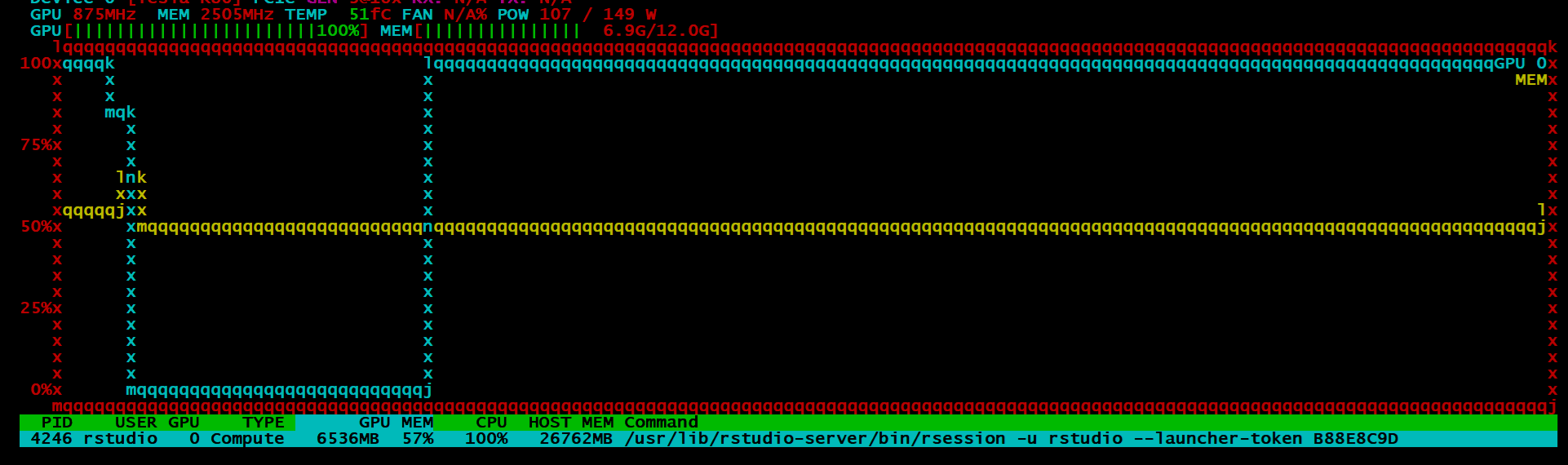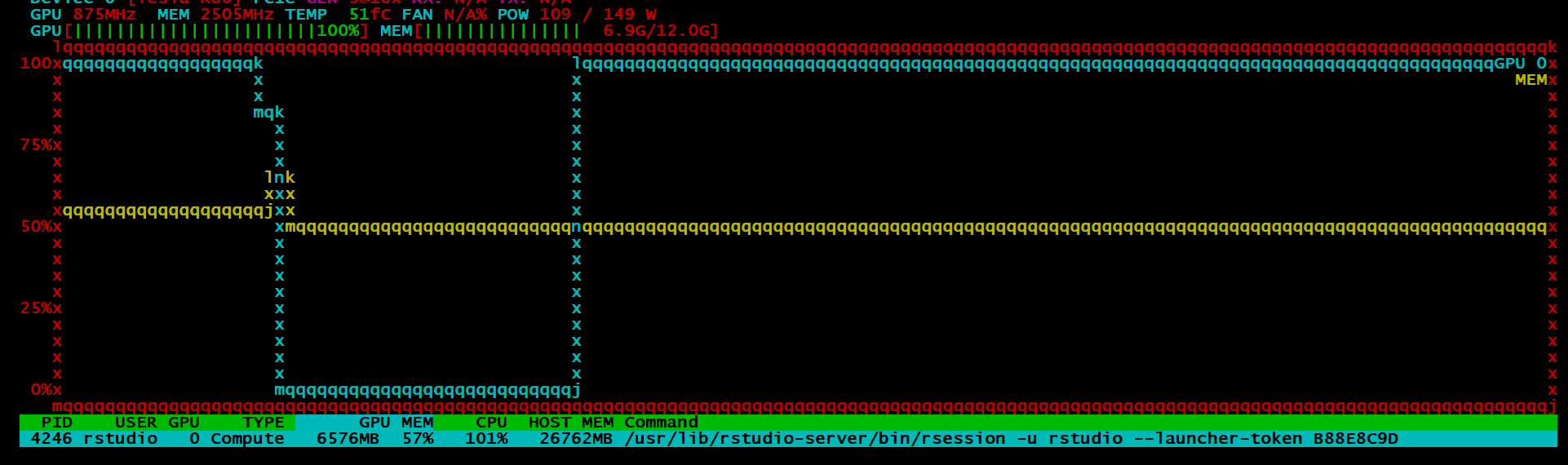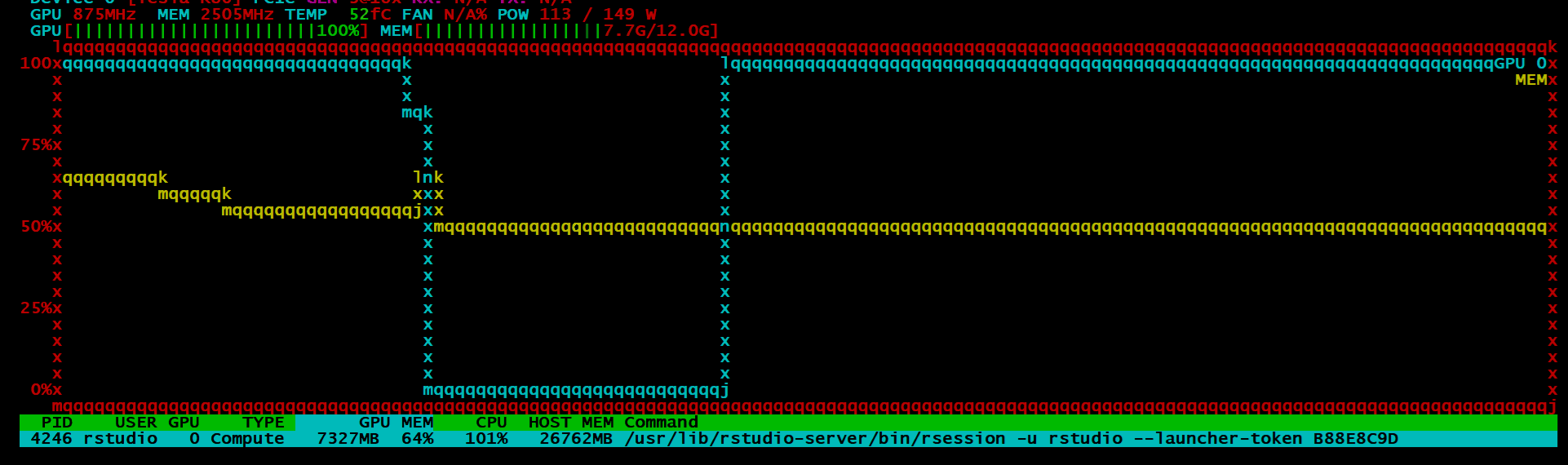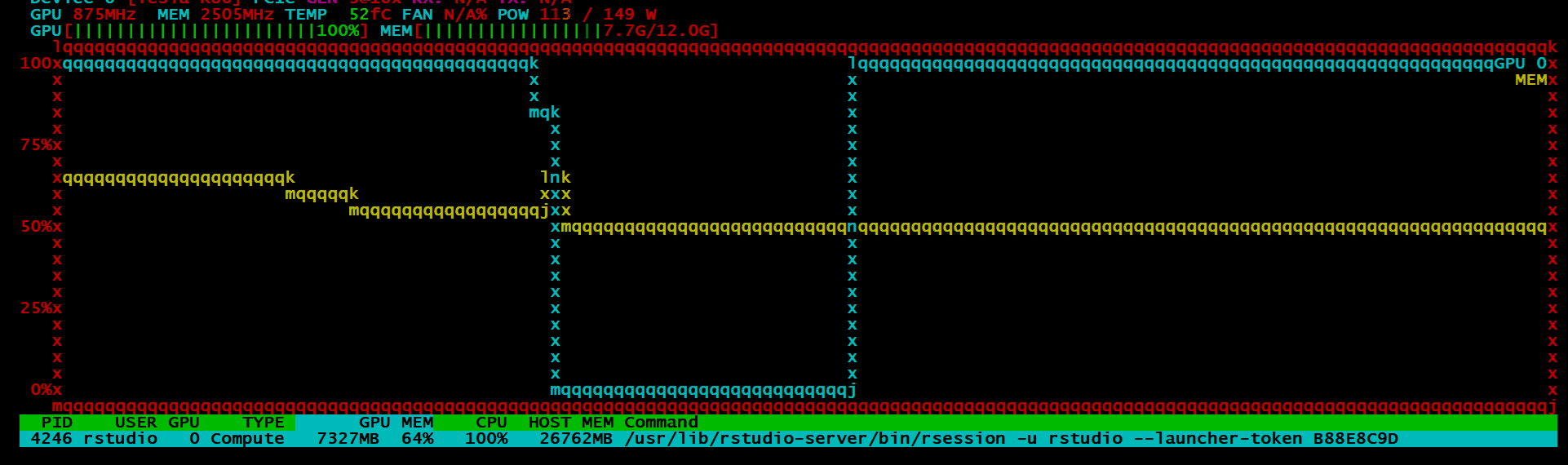
Conclusion
In all honesty, I have been itching to play with GPUs to estimate my models, now I realise the itch was just poison ivy. The setup was frustrating and took me a day to figure out. The interface of the h2o4gpu package is not the same as the CPU h2o variant and I am sceptical about the API doing what it says it is doing - the AUC of the Random Forest and GBM was the same! Which is either a crazy concidence or something is wrong. The last thing that irritated me was the lack of documentation on how to go from training to production using the h2o4gpu package - one of the advantages of using the h2o package is its excellent documentation and its focus on being ‘production ready’.
Despite all of this, I can say, the idea is promising and I’ll be watching the package closely. It is still in its infancy, so one cannot be too judgemental4. Training on ~7.7 million observations with 28 variables only took around a minute which is astonishingly fast. The idea was that I would go on the benchmark the h2o4gpu and h2o package in terms of speed and accuracy, but given my scepticism around the output of the model, I might put that off for another day.
Also, seeing that Colin Fay is coming to SatRday 2020 Johannesburg, I think Ill be better off spending time learning golem, than benchmarking tools likely to change in the future.
- Because if you not doing machine learning on GPUs these days, apparently you cannot claim to be a Data Scientist ^
- I might also be wrong, but the code worked and I trudged forward ^
- https://www.linkedin.com/in/erin-ledell ^
- Plus, lets be honest, this stuff ain’t easy and I can certainly not do better ^
